t(7;8)(q34;p11) TRIM24/FGFR1
2006-02-01 Elena Belloni , Francesco Lo Coco , Pier Giuseppe Pelicci Affiliation1.IFOM, Fondazione Istituto FIRC di Oncologia Molecolare, and IEO, Istituto Europero di Oncologia, Milan, Italy
Clinics and Pathology
Disease
Acute myeloid leucemia (AML) and 8p11 myeloproliferative syndrome (EMS).
Chromosome band 8p11 has been implicated in recurrent chromosome rearrangements associated either with acute myeloid leukemia (AML) or with a peculiar myeloproliferative disorder named 8p11 myeloproliferative syndrome (EMS). The latter is characterized by generalized lymphadenopathy and marked eosinophilia, rapid evolution to AML, and frequent association with non-Hodgkin lymphoma. The FGFR1 gene is constantly involved in EMS, often evolving to AML.
Chromosome band 8p11 has been implicated in recurrent chromosome rearrangements associated either with acute myeloid leukemia (AML) or with a peculiar myeloproliferative disorder named 8p11 myeloproliferative syndrome (EMS). The latter is characterized by generalized lymphadenopathy and marked eosinophilia, rapid evolution to AML, and frequent association with non-Hodgkin lymphoma. The FGFR1 gene is constantly involved in EMS, often evolving to AML.
Phenotype stem cell origin
AML-M4 (EMS)
Clinics
1 case to date, a female patient aged 49 yrs. The differential WBC count suggested a chronic MPD. Examination of a bone marrow smear showed 44% blasts, hypoplasia, and eosinophilia. The immunophenotypic characterization revealed the coexistence of two distinct components, a myelomonocytic part along with a lymphoid population. A diagnosis of AML-M4 was established.
Treatment
Induction therapy was started with daunorubicin, cytosine arabinoside, and etoposide.
Evolution
The patient died of sepsis during aplasia on day 20.
Cytogenetics
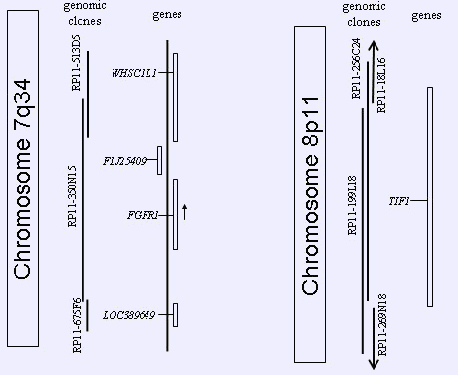
Genomic clones and genes in the FGFR1 (8p11) and TIF1 (7q34) regions. To the left are shown the positions of the RPCI11- 350N15 at 8p11 and of the 2 adjacent genomic clones, RPCI11-513D5 and 675F6 (vertical white bars, genes in the genomic region of interest). On the right are the positions of the 4 clones that span the TIF1 locus (indicated by a vertical white bar) at 7q34 (RPCI11-18L16, 256C24, 199L18, and 269N18).
Additional anomalies
none
Genes Involved and Proteins
Gene name
TRIM24 (tripartite motif-containing 24)
Location
7q33
Note
Alias TIF1
Dna rna description
see figure a. Transcript: 2 variants. For details see the specific NCBI page.
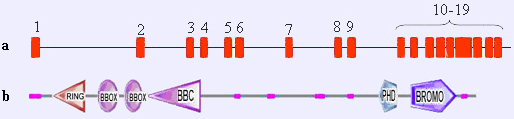
Genomic structure (not drawn to scale) of the TIF1 loci (numbered black boxes, exons) in figure a. The corresponding protein are represented in figure b.
Protein description
TIF1 encodes a nuclear protein, transcription intermediary factor 1a displaying an RBCC motif (RING finger, B-BOX, and coiled-coil domains, also called tripartite motif, TRIM) in its N-terminus and PHD and bromo domains at the C-terminus (figure b).
Gene name
FGFR1 (Fibroblast Growth Factor Receptor 1)
Location
8p11.23
Dna rna description
see figure c. Transcript: various isoforms. For details see the specific NCBI page.
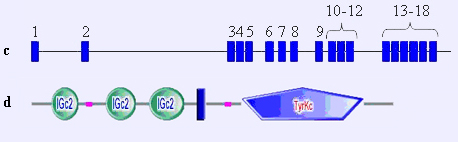
Genomic structure (not drawn to scale) of the FGFR1 loci (numbered black boxes, exons) in figure c. The corresponding proteins are represented in figure d.
Protein description
The FGFR1 gene encodes the fibroblast growth factor receptor 1, a transmembrane receptor with a tyrosine kinase (TK) domain in the intracellular C-terminus, a transmembrane (TM) domain, and 3 immunoglobulin-like C-2 type extracellular domains.
Result of the Chromosomal Anomaly
Note
TIF1-FGFR1 and FGFR1-TIF1: see figure e.
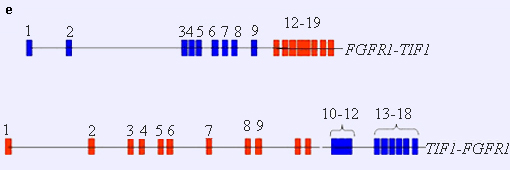
Note
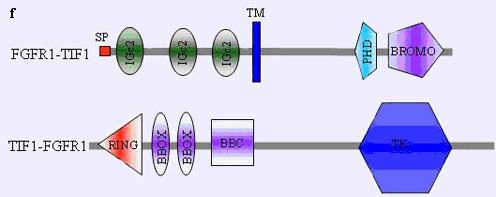
TIF1-FGFR1 and FGFR1-TIF1: see figure f.
Highly cited references
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 36709160 | 2022 | [8p11 myeloproliferative syndrome with TRIM24/FGFR1 and FGFR1/TRIM24 fusion gene positive: a case report]. | 0 |
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 15609342 | 2005 | 8p11 myeloproliferative syndrome with a novel t(7;8) translocation leading to fusion of the FGFR1 and TIF1 genes. | Belloni E et al |
| 11640868 | 2001 | Myeloproliferative disorders. | Bench AJ et al |
| 10640219 | 1998 | Molecular genetics and cytogenetics of myeloproliferative disorders. | Bench AJ et al |
| 15995322 | 2005 | Molecular classification and pathogenesis of eosinophilic disorders: 2005 update. | Gotlib J et al |
| 16781488 | 2006 | Eosinophilic disorders: molecular pathogenesis, new classification, and modern therapy. | Gotlib J et al |
| 15034873 | 2004 | Identification of a novel gene, FGFR1OP2, fused to FGFR1 in 8p11 myeloproliferative syndrome. | Grand EK et al |
| 16946300 | 2006 | Phosphotyrosine profiling identifies the KG-1 cell line as a model for the study of FGFR1 fusions in acute myeloid leukemia. | Gu TL et al |
| 14660670 | 2004 | Critical role of STAT5 activation in transformation mediated by ZNF198-FGFR1. | Heath C et al |
| 9889006 | 1999 | The genomic structure of ZNF198 and location of breakpoints in the t(8;13) myeloproliferative syndrome. | Kulkarni S et al |
| 11919391 | 2002 | The 8p11 myeloproliferative syndrome: a distinct clinical entity caused by constitutive activation of FGFR1. | Macdonald D et al |
| 9716603 | 1998 | Consistent fusion of ZNF198 to the fibroblast growth factor receptor-1 in the t(8;13)(p11;q12) myeloproliferative syndrome. | Reiter A et al |
| 10373756 | 1999 | [The 8p11 myeloproliferative syndrome]. | Reither A et al |
| 15050920 | 2004 | Distinct stem cell myeloproliferative/T lymphoma syndromes induced by ZNF198-FGFR1 and BCR-FGFR1 fusion genes from 8p11 translocations. | Roumiantsev S et al |
| 10935490 | 1999 | ZNF198-FGFR1 transforms Ba/F3 cells to growth factor independence and results in high level tyrosine phosphorylation of STATS 1 and 5. | Smedley D et al |
| 11550283 | 2001 | Identification of four new translocations involving FGFR1 in myeloid disorders. | Sohal J et al |
| 15995325 | 2005 | Modern diagnosis and treatment of primary eosinophilia. | Tefferi A et al |
| 15570299 | 2004 | Clinical variability of patients with the t(6;8)(q27;p12) and FGFR1OP-FGFR1 fusion: two further cases. | Vizmanos JL et al |
| 15800673 | 2005 | The t(8;17)(p11;q23) in the 8p11 myeloproliferative syndrome fuses MYO18A to FGFR1. | Walz C et al |
| 16777224 | 2007 | Clonal evolution of 8p11 stem cell syndrome in a 14-year-old Chinese boy: a review of literature of t(8;13) associated myeloproliferative diseases. | Wong WS et al |
| 16879608 | 2006 | A biphenotypic transformation of 8p11 myeloproliferative syndrome with CEP1/FGFR1 fusion gene. | Yamamoto K et al |
Summary
Fusion gene
TRIM24/FGFR1 TRIM24 (7q33) FGFR1 (8p11.23) M|TRIM24/FGFR1 TRIM24 (7q33) FGFR1 (8p11.23) M t(7;8)(q34;p11)|TRIM24/FGFR1 TRIM24 (7q33) FGFR1 (8p11.23) TIC
Citation
Elena Belloni ; Francesco Lo Coco ; Pier Giuseppe Pelicci
t(7;8)(q34;p11) TRIM24/FGFR1
Atlas Genet Cytogenet Oncol Haematol. 2006-02-01
Online version: http://atlasgeneticsoncology.org/haematological/1409/t(7;8)(q34;p11)-trim24-fgfr1
