CASZ1 (castor zinc finger 1)
2013-01-01 Zhihui Liu , Nancy Ding , Carol J Thiele AffiliationPediatric Oncology Branch, National Cancer Institute, National Institutes of Health, USA
Identity
HGNC
LOCATION
1p36.22
LOCUSID
ALIAS
CAS11,CST,SRG,ZNF693,dJ734G22.1
FUSION GENES
DNA/RNA
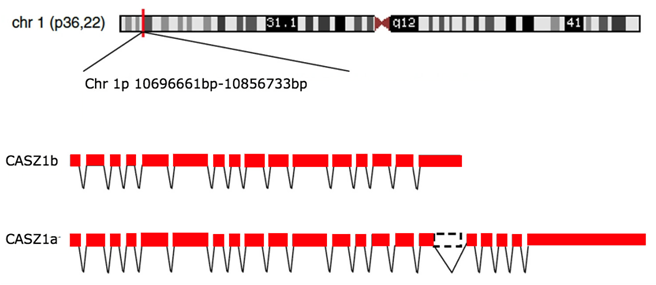
Location of DNA of CASZ1 and two forms of the gene due to alternative splicing.
Description
DNA of CASZ1a contains 160072 bp composed of 21 exons with 11 zinc fingers, alternative splicing at exon 16 of CASZ1a can result in CASZ1b (16 exons with 5 zinc fingers) (Liu et al., 2006).
Transcription
CASZ1 mRNA transcribed in centromeric to telomeric orientation. The CASZ1a cDNA sequence spans 7964 bp, with a 5 UTR of 346 bp, an open reading frame of 5280 bp, and a 3 UTR of 2339 bp; the CASZ1b cDNA sequence spans 4438 bp, with a 5 UTR of 346 bp, an open reading frame of 3501 bp, and a 3 UTR of 592 bp (Liu et al., 2006; with updated information from NCBI database).
Proteins
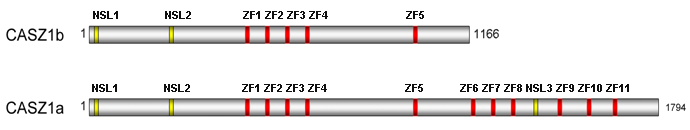

Functional domains of CASZ1.
Description
The 5 of CASZ1 contains 2 nuclear localization signals, which tag proteins for import into the cell nucleus, followed by 5 zinc fingers of C2H2 type (Cys2His2). C2H2 zinc fingers are thought to have DNA binding and protein-protein mediation roles. For CASZ1a, six other zinc fingers (ZF) are present along with a third nuclear localization signal between ZF8 and ZF9. CASZ1b totals 1166 amino acids and CASZ1a totals 1794 amino acids (Liu et al., 2006; Virden et al., 2012).
Expression
Increased expression when cells of neural and mesenchymal origin are induced to differentiation in vitro. CASZ1 is highly expressed in eye, heart, lung, skeletal muscle, pancreas, testis, small intestine and stomach (Liu et al., 2006; Liu et al., 2011a). CASZ1 is dynamically expressed in the development heart and during neurogenesis (Vacalla and Theil, 2002).
Localisation
Nucleus, cytoplasm.
Function
CASZ1 is a crucial zinc finger transcription factor that controls the differentiation of neural (Cui and Doe, 1992) and cardiac muscle cells (Christine and Conlon, 2008), eye development (Vacalla and Theil, 2002) and has tumor suppressor properties (Liu et al., 2011b). There is monoallelic loss of CASZ1 in neuroblastoma (NB) tumors through loss of heterozygosity (LOH) and epigenetic silencing (Wang et al., 2012). Restoration of CASZ1 in NB cells induces cell differentiation, inhibits cell migration and suppresses NB growth (Liu et al., 2011a; Liu et al., 2011b). Structural and functional studies show that either deletion of the N-terminus or mutation in any of the first four zinc fingers of CASZ1b results in a drastic loss in transcriptional activity, which also leads to a decreased tumor suppression activity (Virden et al., 2012). Additionally, single nucleotide polymorphisms in this gene are associated with blood pressure variations in East Asian populations (Takeuchi et al., 2010).
Homology
Unknown.
Mutations
Note
Unknown.
Implicated in
Entity name
Neuroblastoma
Note
Neuroblastomas (NB) are pediatric cancers arising from cells of the sympathoadrenal lineage of neural crest cells (Brodeur, 2003; Maris, 2010). CASZ1 maps to 1p36.22, part of the most commonly deleted regions of chromosome 1p that occur in 35% of NB cases (Brodeur, 2003). A recent study in 184 primary NB showed that 1p36.3 is the shortest region of overlap for loss of heterozygosity (LOH) (White et al., 2005), although 98% of these 1p36.3 LOH NB tumors also have lost the region encompassing CASZ1 (Liu et al., 2011b). Additionally, CASZ1 is epigenetically silenced in NB by EZH2, a methyltransferase (Wang et al., 2012). HDAC inhibitor treatment of NB cells increases CASZ1 expression (Carén et al., 2007; Fransson et al., 2007). Consistent with CASZ1 being epigenetically regulated, a bivalent mark was found in CASZ1 promoter region in ES cells (Bernstein et al., 2006). CASZ1 is upregulated when NB cells are induced to differentiate by retinoic acid, and restoration of CASZ1 in NB cells up-regulates differentiation genes such as NGFR and TrkA, and suppresses tumor growth (Liu et al., 2011a; Liu et al., 2011b). In NB patients, low CASZ1 expression in NB tumors is associated with poor prognosis and an undifferentiated histology, which suggests loss of CASZ1 may contribute to NB tumorigenesis.
Entity name
Cervical carcinoma
Note
Human papillomavirus (HPV) genome integration into host chromatin enables expression of oncogenes E6 and E7 leading to cervical carcinogenesis. HPV integration may affect transcriptionally active sites. In one reported case of cervical cancer, HPV was found to functionally delete CASZ1 (Schmitz et al., 2012).
Entity name
Blood hypertension
Note
Hypertension, increases in blood pressure, can lead to cardiovascular disease including stroke and ischemic heart disease. CASZ1 has been found to be one of seven loci associated with hypertension in East Asian populations (Takeuchi et al., 2010). CASZ1 could be linked to heart malfunction through inflammatory responses. CASZ1 had been found to have associations in European ancestry but Japanese loci effect size was significantly larger (Takeuchi et al., 2010).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 16630819 | 2006 | A bivalent chromatin structure marks key developmental genes in embryonic stem cells. | Bernstein BE et al |
| 12612655 | 2003 | Neuroblastoma: biological insights into a clinical enigma. | Brodeur GM et al |
| 17940511 | 2007 | Genetic and epigenetic changes in the common 1p36 deletion in neuroblastoma tumours. | Carén H et al |
| 18410736 | 2008 | Vertebrate CASTOR is required for differentiation of cardiac precursor cells at the ventral midline. | Christine KS et al |
| 1339340 | 1992 | ming is expressed in neuroblast sublineages and regulates gene expression in the Drosophila central nervous system. | Cui X et al |
| 17044048 | 2007 | Neuroblastoma tumors with favorable and unfavorable outcomes: Significant differences in mRNA expression of genes mapped at 1p36.2. | Fransson S et al |
| 21490919 | 2011 | CASZ1b, the short isoform of CASZ1 gene, coexpresses with CASZ1a during neurogenesis and suppresses neuroblastoma cell growth. | Liu Z et al |
| 21252912 | 2011 | CASZ1, a candidate tumor-suppressor gene, suppresses neuroblastoma tumor growth through reprogramming gene expression. | Liu Z et al |
| 20558371 | 2010 | Recent advances in neuroblastoma. | Maris JM et al |
| 22262398 | 2012 | Loss of gene function as a consequence of human papillomavirus DNA integration. | Schmitz M et al |
| 20479155 | 2010 | Blood pressure and hypertension are associated with 7 loci in the Japanese population. | Takeuchi F et al |
| 12351199 | 2002 | Cst, a novel mouse gene related to Drosophila Castor, exhibits dynamic expression patterns during neurogenesis and heart development. | Vacalla CM et al |
| 22331471 | 2012 | Characterization of critical domains within the tumor suppressor CASZ1 required for transcriptional regulation and growth suppression. | Virden RA et al |
| 22068036 | 2012 | EZH2 Mediates epigenetic silencing of neuroblastoma suppressor genes CASZ1, CLU, RUNX3, and NGFR. | Wang C et al |
| 15829979 | 2005 | Definition and characterization of a region of 1p36.3 consistently deleted in neuroblastoma. | White PS et al |
Other Information
Locus ID:
NCBI: 54897
MIM: 609895
HGNC: 26002
Ensembl: ENSG00000130940
Variants:
dbSNP: 54897
ClinVar: 54897
TCGA: ENSG00000130940
COSMIC: CASZ1
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000130940 | ENST00000344008 | Q86V15 |
| ENSG00000130940 | ENST00000377022 | Q86V15 |
| ENSG00000130940 | ENST00000447850 | K7EQC6 |
Expression (GTEx)
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38058200 | 2024 | CASZ1 upregulates PI3K-AKT-mTOR signaling and promotes T-cell acute lymphoblastic leukemia. | 1 |
| 38058200 | 2024 | CASZ1 upregulates PI3K-AKT-mTOR signaling and promotes T-cell acute lymphoblastic leukemia. | 1 |
| 38053207 | 2023 | Role of the CASZ1 transcription factor in tissue development and disease. | 0 |
| 38053207 | 2023 | Role of the CASZ1 transcription factor in tissue development and disease. | 0 |
| 36243768 | 2022 | Loss of CASZ1 tumor suppressor linked to oncogenic subversion of neuroblastoma core regulatory circuitry. | 4 |
| 36243768 | 2022 | Loss of CASZ1 tumor suppressor linked to oncogenic subversion of neuroblastoma core regulatory circuitry. | 4 |
| 33205602 | 2021 | Circular RNA circANKRD36 regulates Casz1 by targeting miR-599 to prevent osteoarthritis chondrocyte apoptosis and inflammation. | 18 |
| 33205602 | 2021 | Circular RNA circANKRD36 regulates Casz1 by targeting miR-599 to prevent osteoarthritis chondrocyte apoptosis and inflammation. | 18 |
| 32060262 | 2020 | CASZ1 induces skeletal muscle and rhabdomyosarcoma differentiation through a feed-forward loop with MYOD and MYOG. | 25 |
| 32060262 | 2020 | CASZ1 induces skeletal muscle and rhabdomyosarcoma differentiation through a feed-forward loop with MYOD and MYOG. | 25 |
| 31570750 | 2019 | A variant of the castor zinc finger 1 (CASZ1) gene is differentially associated with the clinical classification of chronic venous disease. | 6 |
| 31642367 | 2019 | Epigenome-Wide Association Study for All-Cause Mortality in a Cardiovascular Cohort Identifies Differential Methylation in Castor Zinc Finger 1 (CASZ1). | 14 |
| 31570750 | 2019 | A variant of the castor zinc finger 1 (CASZ1) gene is differentially associated with the clinical classification of chronic venous disease. | 6 |
| 31642367 | 2019 | Epigenome-Wide Association Study for All-Cause Mortality in a Cardiovascular Cohort Identifies Differential Methylation in Castor Zinc Finger 1 (CASZ1). | 14 |
| 29467767 | 2018 | Identification of Casz1 as a Regulatory Protein Controlling T Helper Cell Differentiation, Inflammation, and Immunity. | 15 |
Citation
Zhihui Liu ; Nancy Ding ; Carol J Thiele
CASZ1 (castor zinc finger 1)
Atlas Genet Cytogenet Oncol Haematol. 2013-01-01
Online version: http://atlasgeneticsoncology.org/gene/45989/casz1
