MARK4 (MAP/microtubule affinity-regulating kinase 4)
2003-05-01 Alessandro Beghini, PhD AffiliationUniversity of Milan, Medical Faculty, Department of Biology, Genetics for Medical Sciences, via Viotti 5, 20133-Milan, Italy
Identity
HGNC
LOCATION
19q13.32
IMAGE

LOCUSID
ALIAS
MARK4L,MARK4S,MARKL1,MARKL1L,PAR-1D
FUSION GENES
DNA/RNA
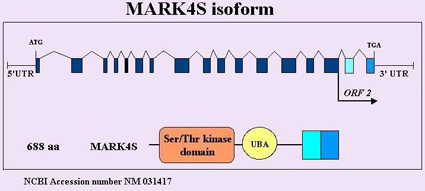
Description
Spans 55,6 kb; 18 exons
Transcription
3,6kb mRNA of MARK4S isoform, 3,22kb of MARK4L isoform (alternative splicing-skipping of exon 16, which leads to a change in the reading frame ).
Proteins
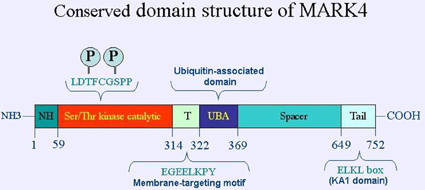
Description
688 amino acids (aa) for MARK4S isoform and 752 aa for MARK4L isoform; belongs to the MARK family of protein kinases and contains from aa 59 to 314 a Serine-Threonine kinase catalytic domain with two activating phosphorylation sites. A short sequence (T region) contains a putative membrane-targeting motif. This region is followed by a ubiquitin-associated (UBA) domain. The spacer is the least-conserved region among MARKs proteins. This region is followed by a strikingly conserved C-terminal domain. In MARK4 the C-terminal domain differs between MARK4S and MARK4L isoforms.
Expression
The MARK4S isoform is predominantly expressed in the brain and at low levels in the heart. The MARK4L isoform is expressed ubiquitously in all tissues, with a highly abundant expression in testis, neural progenitors and glial tumors. MARK4L is downregulated during glial differentiation.
Localisation
Protein was detected homogeneously in cytoplasm.
Function
MARK4 is considered to play a role as a messenger of the Wnt-signaling pathway. MARK4L represents a mitogenic-associated isoform.
Homology
MARK1, MARK2 (Emk1), MARK3 (p78/C-TAK1), par1, kin1
Mutations
Note
Mutations have not been detected.
Implicated in
Entity name
Hepatocellular carcinogenesis
Oncogenesis
RT-PCR anaysis detected upregulated expression in nearly all clinical hepatocellular carcinoma cells in which nuclear accumulation of b-catenin was observed.
Entity name
Up-regulation and overexpression of MARK4 has been described in glial tumors and glioblastoma cell lines.
Oncogenesis
MARK4 gene activation (enhanced expression and/or amplification) may result from intrachromosomal duplication upon 19q rearrangements.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 12735302 | 2003 | The neural progenitor-restricted isoform of the MARK4 gene in 19q13.2 is upregulated in human gliomas and overexpressed in a subset of glioblastoma cell lines. | Beghini A et al |
| 9757832 | 1998 | MAPs, MARKs and microtubule dynamics. | Drewes G et al |
| 11326310 | 2001 | Isolation of a novel human gene, MARKL1, homologous to MARK3 and its involvement in hepatocellular carcinogenesis. | Kato T et al |
| 11347906 | 2001 | Prediction of the coding sequences of unidentified human genes. XX. The complete sequences of 100 new cDNA clones from brain which code for large proteins in vitro. | Nagase T et al |
Other Information
Locus ID:
NCBI: 57787
MIM: 606495
HGNC: 13538
Ensembl: ENSG00000007047
Variants:
dbSNP: 57787
ClinVar: 57787
TCGA: ENSG00000007047
COSMIC: MARK4
RNA/Proteins
Expression (GTEx)
Pathways
| Pathway | Source | External ID |
|---|---|---|
| Organelle biogenesis and maintenance | REACTOME | R-HSA-1852241 |
| Cilium Assembly | REACTOME | R-HSA-5617833 |
| Anchoring of the basal body to the plasma membrane | REACTOME | R-HSA-5620912 |
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38041405 | 2024 | Gain-of-function MARK4 variant associates with pediatric neurodevelopmental disorder and dysmorphism. | 0 |
| 38301049 | 2024 | Interaction of Sp1 and Setd8 promotes vascular smooth muscle cells apoptosis by activating Mark4 in vascular calcification. | 1 |
| 38041405 | 2024 | Gain-of-function MARK4 variant associates with pediatric neurodevelopmental disorder and dysmorphism. | 0 |
| 38301049 | 2024 | Interaction of Sp1 and Setd8 promotes vascular smooth muscle cells apoptosis by activating Mark4 in vascular calcification. | 1 |
| 34751461 | 2022 | Myricetin inhibits breast and lung cancer cells proliferation via inhibiting MARK4. | 11 |
| 35192892 | 2022 | MARK2/4 promotes Warburg effect and cell growth in non-small cell lung carcinoma through the AMPKα1/mTOR/HIF-1α signaling pathway. | 6 |
| 34751461 | 2022 | Myricetin inhibits breast and lung cancer cells proliferation via inhibiting MARK4. | 11 |
| 35192892 | 2022 | MARK2/4 promotes Warburg effect and cell growth in non-small cell lung carcinoma through the AMPKα1/mTOR/HIF-1α signaling pathway. | 6 |
| 32141394 | 2021 | Impact of glioblastoma multiforme associated mutations on the structure and function of MAP/microtubule affinity regulating kinase 4. | 5 |
| 32141394 | 2021 | Impact of glioblastoma multiforme associated mutations on the structure and function of MAP/microtubule affinity regulating kinase 4. | 5 |
| 31411112 | 2020 | Structure and dynamics of inactive and active MARK4: conformational switching through the activation process. | 4 |
| 32535203 | 2020 | Structural and biochemical investigation of MARK4 inhibitory potential of cholic acid: Towards therapeutic implications in neurodegenerative diseases. | 12 |
| 33020179 | 2020 | Microtubule affinity-regulating kinase 4 with an Alzheimer's disease-related mutation promotes tau accumulation and exacerbates neurodegeneration. | 16 |
| 31411112 | 2020 | Structure and dynamics of inactive and active MARK4: conformational switching through the activation process. | 4 |
| 32535203 | 2020 | Structural and biochemical investigation of MARK4 inhibitory potential of cholic acid: Towards therapeutic implications in neurodegenerative diseases. | 12 |
Citation
Alessandro Beghini, PhD
MARK4 (MAP/microtubule affinity-regulating kinase 4)
Atlas Genet Cytogenet Oncol Haematol. 2003-05-01
Online version: http://atlasgeneticsoncology.org/gene/419/mark4
