RBM5 (RNA binding motif protein 5)
2007-02-01 Mirna Mourtada-Maarabouni AffiliationSchool of Life Sciences, Keele University, Keele, Staffs, ST5 5BG, United Kingdom
Identity
HGNC
LOCATION
3p21.31
LOCUSID
ALIAS
G15,H37,LUCA-15,LUCA15,RMB5
FUSION GENES
DNA/RNA
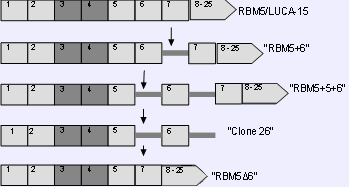
RBM5/LUCA-15 splice variants. Boxes represent the exons, intronic sequences are represented by horizontal lines. The arrows point to the STOP codon in the protein coding sequence.
Description
The gene spans about 30.03 Kb. Orientation plus strand. At least 25 exons.
Transcription
Full length RBM5 has 3.1 Kb, 2448 bp open reading frame. The gene encodes a number of alternative splice variants, identified by reverse transcription polymerase chain reaction. One splice variant lacks exon 6, RBM5delta6. Three other RNA splice variants retain intronic sequences, RBM5+5+6 retains introns 5 and 6, RBM5+6 retains intron 6 and clone 26 which is a partial cDNA containing an open reading frame and terminates within intron 6. A 326 bp antisense cDNA that maps to the intronic region of RBM5, je2, has also been identified. Both, RBM5delta6 and clone 26 cDNAs have been cloned.
Proteins
Description
Full length RBM5 encodes a protein with a molecular mass of about 90 kDa (815 amino acids). The protein has two RNA Binding Domains (RBD), also recognized as RNA Recognition Motif (RRM). RBM5 structure also features other functional motifs, which includes two putative zinc-finger DNA binding motifs, two bipartite nuclear localisation signals and a G-patch domain (a conserved domain in eukaryotic RNA-processing proteins and type D retroviral polyproteins), suggesting a role for RBM5/LUCA-15 in RNA processing. In addition, the C-terminal region of RBM5/LUCA-15 contains several domains including a glutamine rich domain (362-385), which is thought to serve as protein-protein interaction site in certain RNA- and DNA- binding proteins.
RBM5delta6 encodes a protein of 17 kDa, due to a frameshift mutation caused by the deletion of exon 6. The only functional motif retained by this truncated protein is the arginine/glycine-rich amino terminal region. A putative 21 kDa is reported to be encoded by RBM5+5+6 and clone 26, as revealed by an in vitro transcription/translation study in rabbit reticulocyte lysate.
RBM5delta6 encodes a protein of 17 kDa, due to a frameshift mutation caused by the deletion of exon 6. The only functional motif retained by this truncated protein is the arginine/glycine-rich amino terminal region. A putative 21 kDa is reported to be encoded by RBM5+5+6 and clone 26, as revealed by an in vitro transcription/translation study in rabbit reticulocyte lysate.
Expression
Full length RBM5 and its alternative splice variants RNA are widely expressed in both primary tissue and cell lines. Northern blot analysis revealed higher expression in skeletal and heart muscles and pancreas. The expression of the full length RBM5 mRNA is reported to be high in adult thymus and low in the foetal thymus, suggesting that its expression is developmentally regulated. Full length RBM5/LUCA-15 is reported to be down-regulated in breast cancer specimens, in 82% of primary non-small-cell lung carcinoma specimens and in many lung cancer cell lines and in vestibular schwannomas (27 fold reduction). The expression of RBM5delta6 RNA is highest in transformed cells. The full length RBM5 protein is ubiquitously expressed in human tissues. It was found to be downregulated in 73% of primary non-small-cell lung carcinoma specimens compared to normal adjacent tissue. RBM5/LUCA-15 was one of the antigens identified by autologous antibody in patients with renal carcinoma. Overexpression of je2, the 326 bp antisense sequence, resulted in the downregulation of the RBM5 protein (95 kDa) and the upregulation of the 17 kDa protein (RBM5delta6).
Localisation
RBM5 protein includes bipartite nuclear localisation signals suggesting its localisation to the nucleus.
Function
RBM5/LUCA-15 is among the 35 genes located within the 370 kb overlapping lung cancer homozygous deletion region at 3p21.3. RBM5 has been implicated in the control of cell death by apoptosis and cell proliferation. RBM5s involvement in apoptosis and malignancy has been the focus of many recent studies, with all results converging on a role for RBM5/LUCA-15 as a Tumor Suppressor Gene (TSG). RBM5 splice variants have been shown to function as regulators of apoptosis. Stable expression of Je2, in human T-cell lines CEM-C7 and Jurkat produced marked inhibition of Fas-mediated apoptosis and conferred protection from apoptosis induced by other stimuli. RBM5delta6 inhibits CD95-mediated apoptosis as well as accelerating cell proliferation.
Overexpression of the full length RBM5 in CEM-C7 and Jurkat T-cell lines suppressed cell proliferation both by inducing apoptosis and by inducing cell cycle arrest in G1. Consistently, RBM5/LUCA-15 overexpression also inhibited human lung cancer cell growth (A549 non-small cell lung cancer line) by increasing apoptosis and inducing cell cycle arrest in G1. This inhibition of cell growth was reported to be associated with decreased cyclin A and phosphorylated RB and an increase in the level of the proapototic protein Bax. Introducing RBM5/LUCA-15 into breast cancer cells that had 3p21-22 deletions reduced both anchorage-dependent and anchorage-independent growth. RBM5/LUCA-15 also is reported to suppress anchorage-dependent and anchorage-independent growth in A9 mouse fibrosarcoma cells and to inhibit their tumour forming activity in nude mice. RBM5/LUCA-15 was one of the nine genes down-regulated in metastasis and it has been included in the 17 common gene signature associated with metastasis identified in multiple solid tumour types. Solid tumours carrying this gene expression signature had high rates of metastasis and poor clinical outcome. Microarray-based analysis of changes in gene expression caused by the modulation of the level of RBM5/LUCA-15 were carried out. Among 5603 genes on cDNA microarray, 35 genes, well known for their roles in the cell cycle as well as in apoptosis, were found to be differentially expressed as a result of RBM5/LUCA-15 overexpression in CEM-C7 cells.
These RBM5/LUCA-15 modulated genes include a number of cyclin-dependent kinase complexes, most notably CDK2 and three putative oncogenes which are down-regulated by RBM5/LUCA-15. These are Pim-1, a serine/threonine kinase, ITPA (inosine triphosphate pyrophosphatase) and Amplified in Breast cancer 1 (AIB1). Other functionally important genes modulated by RBM5/LUCA-15 are well established to play specific anti apoptotic roles such as BIRC4 (AIP3), BIRC3 (cIAP2, MIHC), Mcl-1, a member of the Bcl-2 family of apoptosis suppressors, and TRAF1, supporting a role for RBM5/LUCA-15 in apoptosis. Other genes upregulated by RBM5/LUCA-15 include Stat5b, Annexin I and Bone Morphogenetic Protein 5 (BMP5), implicated in apoptosis, immunosuppression and organ development respectively.
Overexpression of the full length RBM5 in CEM-C7 and Jurkat T-cell lines suppressed cell proliferation both by inducing apoptosis and by inducing cell cycle arrest in G1. Consistently, RBM5/LUCA-15 overexpression also inhibited human lung cancer cell growth (A549 non-small cell lung cancer line) by increasing apoptosis and inducing cell cycle arrest in G1. This inhibition of cell growth was reported to be associated with decreased cyclin A and phosphorylated RB and an increase in the level of the proapototic protein Bax. Introducing RBM5/LUCA-15 into breast cancer cells that had 3p21-22 deletions reduced both anchorage-dependent and anchorage-independent growth. RBM5/LUCA-15 also is reported to suppress anchorage-dependent and anchorage-independent growth in A9 mouse fibrosarcoma cells and to inhibit their tumour forming activity in nude mice. RBM5/LUCA-15 was one of the nine genes down-regulated in metastasis and it has been included in the 17 common gene signature associated with metastasis identified in multiple solid tumour types. Solid tumours carrying this gene expression signature had high rates of metastasis and poor clinical outcome. Microarray-based analysis of changes in gene expression caused by the modulation of the level of RBM5/LUCA-15 were carried out. Among 5603 genes on cDNA microarray, 35 genes, well known for their roles in the cell cycle as well as in apoptosis, were found to be differentially expressed as a result of RBM5/LUCA-15 overexpression in CEM-C7 cells.
These RBM5/LUCA-15 modulated genes include a number of cyclin-dependent kinase complexes, most notably CDK2 and three putative oncogenes which are down-regulated by RBM5/LUCA-15. These are Pim-1, a serine/threonine kinase, ITPA (inosine triphosphate pyrophosphatase) and Amplified in Breast cancer 1 (AIB1). Other functionally important genes modulated by RBM5/LUCA-15 are well established to play specific anti apoptotic roles such as BIRC4 (AIP3), BIRC3 (cIAP2, MIHC), Mcl-1, a member of the Bcl-2 family of apoptosis suppressors, and TRAF1, supporting a role for RBM5/LUCA-15 in apoptosis. Other genes upregulated by RBM5/LUCA-15 include Stat5b, Annexin I and Bone Morphogenetic Protein 5 (BMP5), implicated in apoptosis, immunosuppression and organ development respectively.
Homology
RBM5 shares 30% amino acid sequence homology with its immediate telomeric neighbor, the putative tumor suppressor gene RBM6. Another RNA Binding Motif protein that shares high homology with RBM5 (53% at the amino acid level) is RBM10. The RBM10 gene is located on the X chromosome at p11.23. No function has yet been determined for RBM10. The mouse RBM5 gene is 90% identical on cDNA level and 97% on protein level. The fly proteome contain three similar proteins, of which CG4887 gene product shows 43% similarity. The rat binding protein S1-1 has similar domain structure and is close in amino acid sequence to RBM5.
Mutations
Note
Mutations in the RMB5 gene have not been found in lung cancer. RBM5 is reported to be downregulated in RAS-transformed Rat-1 cells, in samples from breast cancer, in human schwannomas and in about 75% of primary lung cancers specimens. RBM5 was one of the nine genes down-regulated in metastasis in primary tumors and it has been included in the 17 common gene signature associated with metastasis identified in multiple solid tumour types. Solid tumours carrying this gene expression signature had high rates of metastasis and poor clinical outcomes.
Implicated in
Entity name
Lung cancer and many other cancers
Disease
RBM5 is located at the human chromosomal locus 3p21.3, which is strongly associated with lung cancer and many other cancers, including head and neck, renal, breast and female genital tract. More than 90% of freshly microdissected primary lung cancers display loss of heterozygosity (LOH) of 3p21.3. RBM5 is implicated in apoptosis and in cell cycle regulation.
Oncogenesis
The ability of RBM5/LUCA-15 to inhibit the growth of transformed cells fulfils one of the criteria for a tumor suppressor. The simultaneous changes in proliferation and inhibition of apoptosis brought about by dysregulation of the RBM5/LUCA-15 locus are likely to be of major significance in oncogenesis.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 10470032 | 1999 | G-patch: a new conserved domain in eukaryotic RNA-processing proteins and type D retroviral polyproteins. | Aravind L et al |
| 8036511 | 1994 | Conserved structures and diversity of functions of RNA-binding proteins. | Burd CG et al |
| 10213508 | 1999 | Molecular cloning of a candidate tumor suppressor gene, DLC1, from chromosome 3p21.3. | Daigo Y et al |
| 8390035 | 1993 | A homozygous deletion on chromosome 3 in a small cell lung cancer cell line correlates with a region of tumor suppressor activity. | Daly MC et al |
| 10353602 | 1999 | DEF-3(g16/NY-LU-12), an RNA binding protein from the 3p21.3 homozygous deletion region in SCLC. | Drabkin HA et al |
| 11029660 | 2000 | LUCA15, a putative tumour suppressor gene encoding an RNA-binding nuclear protein, is down-regulated in ras-transformed Rat-1 cells. | Edamatsu H et al |
| 9500467 | 1998 | Human lung cancer antigens recognized by autologous antibodies: definition of a novel cDNA derived from the tumor suppressor gene locus on chromosome 3p21.3. | Güre AO et al |
| 9365829 | 1997 | Differential elimination of 3p and retention of 3q segments in human/mouse microcell hybrids during tumor growth. | Imreh S et al |
| 7621411 | 1995 | A group of NotI jumping and linking clones cover 2.5 Mb in the 3p21-p22 region suspected to contain a tumor suppressor gene. | Kashuba VI et al |
| 9111863 | 1997 | Deletions of the short arm of chromosome 3 in solid tumors and the search for suppressor genes. | Kok K et al |
| 11085536 | 2000 | The 630-kb lung cancer homozygous deletion region on human chromosome 3p21.3: identification and evaluation of the resident candidate tumor suppressor genes. The International Lung Cancer Chromosome 3p21.3 Tumor Suppressor Gene Consortium. | Lerman MI et al |
| 17195868 | 2006 | The antiapoptotic RBM5/LUCA-15/H37 gene and its role in apoptosis and human cancer: research update. | Maarabouni MM et al |
| 16546166 | 2006 | Candidate tumor suppressor LUCA-15/RBM5/H37 modulates expression of apoptosis and cell cycle genes. | Mourtada-Maarabouni M et al |
| 16585163 | 2006 | 3p21.3 tumor suppressor gene H37/Luca15/RBM5 inhibits growth of human lung cancer cells through cell cycle arrest and apoptosis. | Oh JJ et al |
| 12036935 | 2002 | A candidate tumor suppressor gene, H37, from the human lung cancer tumor suppressor locus 3p21.3. | Oh JJ et al |
| 12469122 | 2003 | A molecular signature of metastasis in primary solid tumors. | Ramaswamy S et al |
| 10508479 | 1999 | Antigens recognized by autologous antibody in patients with renal-cell carcinoma. | Scanlan MJ et al |
| 15514923 | 2005 | RNA binding motif (RBM) proteins: a novel family of apoptosis modulators? | Sutherland LC et al |
| 10352938 | 1999 | A comparison of genomic structures and expression patterns of two closely related flanking genes in a critical lung cancer region at 3p21.3. | Timmer T et al |
| 8603390 | 1996 | Construction of a 600-kilobase cosmid clone contig and generation of a transcriptional map surrounding the lung cancer tumor suppressor gene (TSG) locus on human chromosome 3p21.3: progress toward the isolation of a lung cancer TSG. | Wei MH et al |
| 12218628 | 2002 | cDNA microarray analysis of vestibular schwannomas. | Welling DB et al |
Other Information
Locus ID:
NCBI: 10181
MIM: 606884
HGNC: 9902
Ensembl: ENSG00000003756
Variants:
dbSNP: 10181
ClinVar: 10181
TCGA: ENSG00000003756
COSMIC: RBM5
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38216972 | 2024 | RNA-binding protein RBM5 plays an essential role in acute myeloid leukemia by activating the oncogenic protein HOXA9. | 0 |
| 38272235 | 2024 | CHIP suppresses the proliferation and migration of A549 cells by mediating the ubiquitination of eIF2α and upregulation of tumor suppressor RBM5. | 0 |
| 38619681 | 2024 | ZNF554 Inhibits Endometrial Cancer Progression via Regulating RBM5 and Inactivating WNT/β-Catenin Signaling Pathway. | 0 |
| 38216972 | 2024 | RNA-binding protein RBM5 plays an essential role in acute myeloid leukemia by activating the oncogenic protein HOXA9. | 0 |
| 38272235 | 2024 | CHIP suppresses the proliferation and migration of A549 cells by mediating the ubiquitination of eIF2α and upregulation of tumor suppressor RBM5. | 0 |
| 38619681 | 2024 | ZNF554 Inhibits Endometrial Cancer Progression via Regulating RBM5 and Inactivating WNT/β-Catenin Signaling Pathway. | 0 |
| 36536516 | 2023 | RNA-binding protein PUM2 promotes T-cell acute lymphoblastic leukemia via competitively binding to RBM5 3'UTR with miR-28-5p. | 0 |
| 38031049 | 2023 | Elucidating the role of RBM5 in osteoclastogenesis: a novel potential therapeutic target for osteoporosis. | 0 |
| 36536516 | 2023 | RNA-binding protein PUM2 promotes T-cell acute lymphoblastic leukemia via competitively binding to RBM5 3'UTR with miR-28-5p. | 0 |
| 38031049 | 2023 | Elucidating the role of RBM5 in osteoclastogenesis: a novel potential therapeutic target for osteoporosis. | 0 |
| 35013161 | 2022 | A de novo paradigm for male infertility. | 25 |
| 36535993 | 2022 | A bacterial virulence factor interacts with the splicing factor RBM5 and stimulates formation of nuclear RBM5 granules. | 2 |
| 35013161 | 2022 | A de novo paradigm for male infertility. | 25 |
| 36535993 | 2022 | A bacterial virulence factor interacts with the splicing factor RBM5 and stimulates formation of nuclear RBM5 granules. | 2 |
| 31726810 | 2021 | Circular circPSMC3 inhibits hepatocellular carcinoma migration and invasion by upregulating RBM5. | 1 |
Citation
Mirna Mourtada-Maarabouni
RBM5 (RNA binding motif protein 5)
Atlas Genet Cytogenet Oncol Haematol. 2007-02-01
Online version: http://atlasgeneticsoncology.org/gene/42069/rbm5
