Soft Tissues: Extraskeletal myxoid chondrosarcoma
2017-03-01 Fei Dong , Vickie Jo Affiliation1.Department of Pathology, Brigham and Womens Hospital, Boston, MA
2.Department of Pathology & Oncology, School of Medicine, University of Occupational & Environmental Health, Japan; [email protected]
3.Department of Pathology, Brigham and Womens Hospital, Boston, MA
Abstract
Review on Extraskeletal myxoid chondrosarcoma, with data on clinics, and the genes involved.
Classification
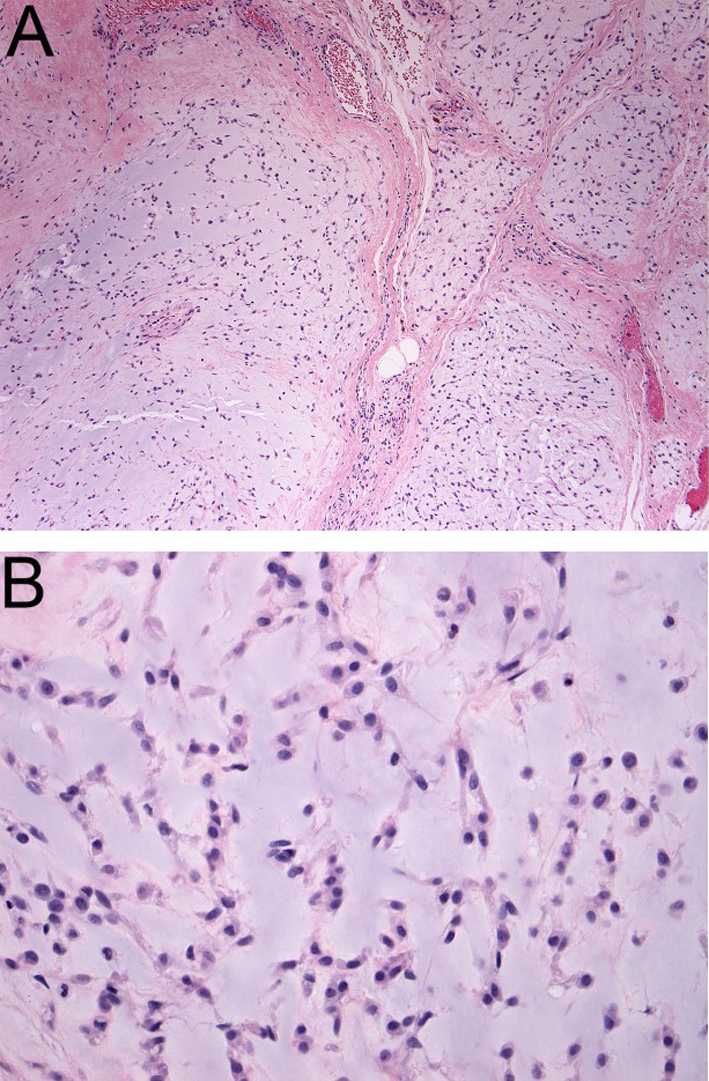
Clinics and Pathology
Epidemiology
Clinics
Pathology
Treatment
Prognosis
Cytogenetics
Note
t(9;22)(q22;q12) EWSR1/NR4A3
t(3;9)(q12;q22) TFG/NR4A3
t(9;11)(q22;q24) HSPA8/NR4A3
t(9;15)(q22;q21) TCF12/NR4A3
t(9;16)(q22;p11) FUS/NR4A3
t(9;17)(q22;q11) TAF15/NR4A3
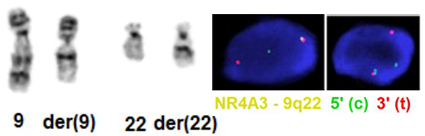
Cytogenetics morphological
Genes Involved and Proteins
Gene name
Location
Dna rna description
Protein description
Gene name
Location
Dna rna description
Protein description
Gene name
Location
Dna rna description
Protein description
Gene name
Location
Dna rna description
Protein description
Gene name
Location
Dna rna description
Protein description
Gene name
Location
Dna rna description
Protein description
Gene name
Location
Dna rna description
Protein description
Gene name
Location
Dna rna description
Protein description
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 24746215 | 2014 | Extraskeletal myxoid chondrosarcoma with non-EWSR1-NR4A3 variant fusions correlate with rhabdoid phenotype and high-grade morphology. | Agaram NP et al |
| 10602520 | 1999 | Identification of a novel fusion gene involving hTAFII68 and CHN from a t(9;17)(q22;q11.2) translocation in an extraskeletal myxoid chondrosarcoma. | Attwooll C et al |
| 24508382 | 2014 | Diagnostic utility of molecular investigation in extraskeletal myxoid chondrosarcoma. | Benini S et al |
| 25130955 | 2014 | Extraskeletal myxoid chondrosarcoma with a t(9;16)(q22;p11.2) resulting in a NR4A3-FUS fusion. | Broehm CJ et al |
| 8570200 | 1996 | Fusion of the EWS gene to CHN, a member of the steroid/thyroid receptor gene superfamily, in a human myxoid chondrosarcoma. | Clark J et al |
| 3967207 | 1985 | Myxoid chondrosarcoma with a translocation involving chromosomes 9 and 22. | Hinrichs SH et al |
| 15188455 | 2004 | TFG is a novel fusion partner of NOR1 in extraskeletal myxoid chondrosarcoma. | Hisaoka M et al |
| 18580682 | 2008 | SMARCB1/INI1 protein expression in round cell soft tissue sarcomas associated with chromosomal translocations involving EWS: a special reference to SMARCB1/INI1 negative variant extraskeletal myxoid chondrosarcoma. | Kohashi K et al |
| 8634690 | 1995 | Oncogenic conversion of a novel orphan nuclear receptor by chromosome translocation. | Labelle Y et al |
| 11156374 | 2000 | Fusion of the NH2-terminal domain of the basic helix-loop-helix protein TCF12 to TEC in extraskeletal myxoid chondrosarcoma with translocation t(9;15)(q22;q21). | Sjögren H et al |
| 28383167 | 2017 | HSPA8 as a novel fusion partner of NR4A3 in extraskeletal myxoid chondrosarcoma. | Urbini M et al |
Citation
Fei Dong ; Vickie Jo
Soft Tissues: Extraskeletal myxoid chondrosarcoma
Atlas Genet Cytogenet Oncol Haematol. 2017-03-01
Online version: http://atlasgeneticsoncology.org/solid-tumor/5025/soft-tissues-extraskeletal-myxoid-chondrosarcoma
Historical Card
2004-09-01 Soft Tissues: Extraskeletal myxoid chondrosarcoma by Masanori Hisaoka,Hiroshi Hashimoto Affiliation
2000-07-01 Soft Tissues: Extraskeletal myxoid chondrosarcoma by Jérôme Couturier Affiliation
